![[Previous Document]](http:/Images/Left.gif)
![[Back to CSI Home Page]](http:/Images/Home.gif)
Below we show an interesting example of MLAB modeling for dimer kinetics. Suppose we have two substances, A and B which bind to form a complex C, and the substance C, in turn, binds with itself to form a dimer D. We thus have:
A + B <--> C with association constant k1 and disassociation constant k2, and
C + C <--> D with association constant k3 and disassociation constant k4.
Suppose further we mix 2 mmoles of A and 3 mmoles of B and measure the concentration in mmoles of both C and D at ten equally-spaced times between 0 and 70 seconds. From this data we wish to estimate the association and dissociation constants k1, k2, k3, and k4. We may proceed in MLAB as follows.
First we read in the data consisting of values of C and D given at the common times 0:70!10. Although common times are used here, this is not required.
* data = (0:70!10) &' read(ddata,10,2)
* type data
time c d
1: 7 1.065 0.0058
2: 14 1.383 0.2203
3: 21 0.9793 0.4019
4: 28 1.107 0.3638
5: 35 0.7289 0.456
6: 42 0.7236 0.5014
7: 49 0.4674 0.715
8: 56 0.6031 0.4723
9: 63 0.6149 0.7219
10: 70 0.3369 0.7294
* cdata = data col 1:2
* ddata = data col (1,3)
Now we define our kinetic model so that c(t) is the concentration of C in mmoles at time t and d(t) is the concentration of D in mmoles at time t.
* fct c't(t)=k1*(a0-c-2*d)*(b0-c-2*d)-k2*c-2*d't(t) * fct d't(t)=k3*c*c-k4*d * initial c(0)=0 * initial d(0)=0 * a0=2;b0=3
Now we guess the values of k1, k2, k3, and k4. We may use the results of equilibrium studies, analyzed by MLAB, to know values for the ratios k1/k2 and k3/k4.
* k1=.02;k2=.002; k3=.02;k4=.002
* constraints q={k1>0, k2>0, k3>0, k4>0}
Now we may curve-fit the two ode-system-defined functions, c and d, to estimate k1, k2, k3, and k4.
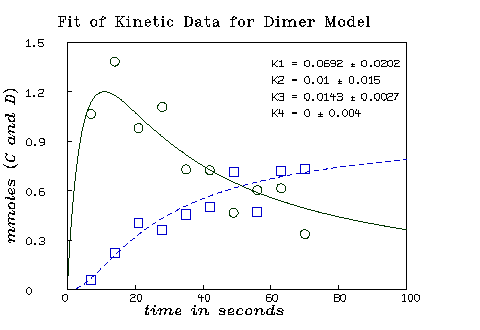
* fit(k1,k2,k3,k4), c to cdata, d to ddata, constraints q final parameter values value error dependency parameter 0.06916895493 0.02023902877 0.5830898606 K1 0.01003638127 0.01500173275 0.5933235047 K2 0.01427015399 0.002713671566 0.7642104538 K3 3.458442141e-20 0.003966026698 0.7396366489 K4 4 iterations CONVERGED best weighted sum of squares = 2.184931e-01 weighted root mean square error = 1.168581e-01 weighted deviation fraction = 1.207618e-01 lagrange multiplier[4] = -3.832523108
Now we may draw the results of the curve-fit.
* m=integrate(c't,d't,0:100!140) * draw m col (1,2) color red * draw m col (1,4) color green lt dashed * draw cdata pt circle lt none color red * draw ddata pt circle lt none color green * bottom title "time in seconds" * left title "mmoles (C and D)" * view
![[Previous Document]](http:/Images/Left.gif)
![[Back to CSI Home Page]](http:/Images/Home.gif)